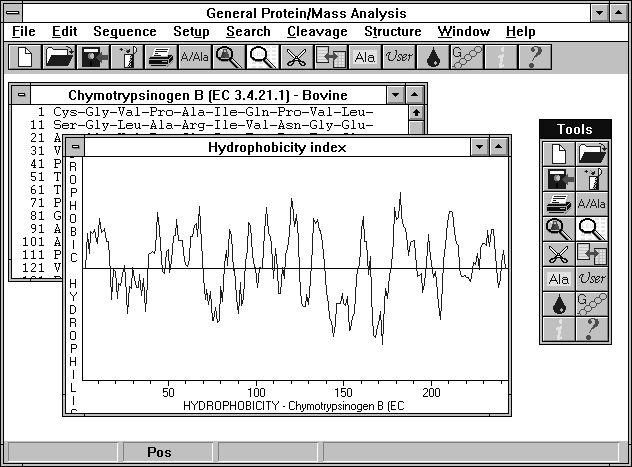
GPMAW was designed specifically for protein chemists for analyzing protein sequences. Protein sequences can easily be entered in the built-in editor or imported from other programs (it interfaces directly to EMBL, European Molecular Biology Laboratory Swiss-Prot Database on-line or on CD ROM-not included). Several sequences can be worked on simultaneously, limited only by computer memory. Cleavage of proteins can be simulated using various enzymes (the list can be edited by the user) and information can be obtained on several parameters; mass (average/monoisotopic), HPLC Index (including graphical representation of separation), Bull & Breeze Index, etc. Proteins can be searched for the presence of peptides of a given mass or composition. Multiple chains, peptide cross-links and modifications of individual amino acid residues can be defined. The program features context sensitive on-line help and a toolbox for easy access to the most common functions. This easy to use, yet powerful program is a must for simulating protein chemistry and can save hours of experimental time. (Requires Windows 3.1 or 95, 98 or NT).
