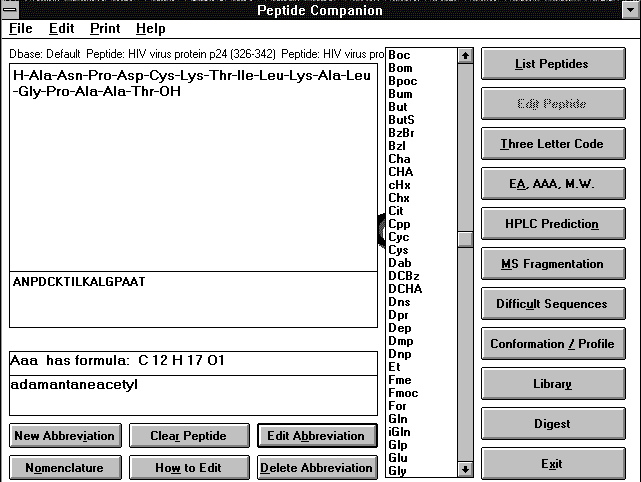
This is a full-featured program designed for chemists, that are working with peptides, peptidometrics, and proteins. It compliments GPMAW (see above). It is capable of handling polynucleotides, carbohydrates, and any organic molecules composed of defined subunits. It calculates molecular weights and results of elemental analysis (which can be corrected for various additional molecules present in lyophilized products) with the option to calculate the analytical result for water and acetic acid content in the product. It evaluates amino acid composition and predicts mass spectral fragmentation of a given peptide or peptidomimetric. The program can calculate the composition of the peptidic or nonpeptidic library, and predicts amino acid composition of a peptide of a given molecular weight. The results of a chemical or enzymatic digestion of a peptide or protein can be calculated and a reversed phase HPLC chromatogram of the digest can be simulated. It can also predict conformational parameters and protein profiles including hydrophobicity, hydrophilicty, and amphipathicity of peptides of up to 3,000 amino acid residues. Standard NIH format for sequence storage is used and therefore sequences from various databases can be imported. Peptide Companion can analyze mass spectra of peptide products and find possible source of problems. It can serve as a database of hundreds of peptide structures and protecting groups. The program uses standard peptide language and it automatically translates three-letter code to one letter code and vice versa. (Requires Windows 3.1, 95, 98 or NT).
